Basic text search
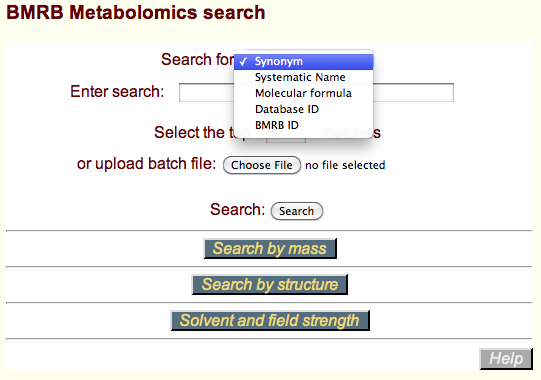
You can use the basic text search for possible names and database IDs. The choice, Synonym, is self evident. The entered term is considered to be a potential substring of all the synonyms in the BMRB metabolomics database. The search engine returns a list of possible matches, ordered by their similarity original term. Systematic name is either a Bielstein standard name or an IUPAC (International Union of Pure and Applied Chemistry) name. For example, caffeine is 1,3,7-trimethylpurine-2,6-quinone.
You can upload a file containing a new line delineated list of synonyms, systematic names, or formulae. If there is already a term in the search field, the engine will append the list to the one in the field. Batch searches for database IDs are more flexible since they do not contain spaces or commas.
Formula: Enter a formula, don't worry about standardization.
Database IDs are usually number based, often with a database identifier. The more specific you can make this search, the more useful it is. Database ID's from PubChem, BMRB, KEGG, ZINC, CAS, MDL and ChEBI can be found here.
Alternatively, you can enter a list of BMRB IDs separated by whitespace or commas.
The search limit will be ignored, when querying the databases or BMRB IDs. This limit would just be annoying.
Databases and BMRB ids can be searched with a comma or space delineated list, either uploaded in a file or entered in the search box. We can't do this with synonyms because it is possible for them to include spaces and other punctuation.
Mass search

Unlike the other mass searches on the BMRB Metabolomics website, which search potential formulae or known metabolites that may or may not be in the BMRB archive, this searches for BMRB entries only. You can select either the standard formula weight or one of four common isotopic labeling schemes for mass spectrometry of biomolecules and NMR spectrometry: natural composition, and 13C, 15N or 13C/15N labeled. Tolerance is defined in either parts per million (ppm) or daltons.
As with the basic text search, you can enter one or more space or comma separated mass values into the text field or upload a file of space, comma, and/or new line separated values for a batch search. If you enter values into the text box and upload a file, the contents of both will be concatenated together and all masses will be queried. If more than one isotopic labeling scheme is chosen, the matching mass will be displayed with bold text.
You can also upload an output file from a Mass Spec experiment as long as it contains a header row with an Mz column to tell the software where the masses are. You are able to account for ionization an common adducts and fragments in the pull-down menu.
Structure search
The structure search is based on standard text representations of molecular connections. The two prevalent chemical graph representations are the simplified molecular-input line-entry specification (SMILES) and International Chemical Identifier (InChI™) strings. Since the InChI string is more likely to unique, it is more likely to yield a valid result.
1D search
Enter either a 1H or 13C space or newline delineated peak list into the text box or upload a newline delineated text file. You can also upload a NMRPipe formatted input file. If you have another file format that you would like to upload, please send us an example of the file you would like to upload to help@bmrb.io.
Because peak searches are meant to determine the possible components of a mixture, we sort the results according to the ratio of number of peaks a metabolite has to how many those peaks matched the search. This can lead to unexpected results sometimes. For instance, if you search against a peak list of pure glucose with a tolerance of 0.03 ppm, methanol is a close match since it only has one peak and it it matches one of the glucose peaks.
2D HSQC peak search
Enter a two columned list of NMR HSQC peaks or upload an NMRPipe or SPARKEY formatted peak list.
Solvent and Field strength
At the moment, this offers the user the ability to search for entries taken in one or more particular solvents, at one or more particular field strengths, or a combinations of the two. This was developed because of user feedback from the macromoleculer side of BMRB. We hope to use this more fully when we incorporate searches on peaks and transitions.
